April 29th, 2022
Brooke Andel and Spencer Harstad
Introduction:
Over the last few weeks, we have spent in the lab, we have continued our work on RNA isolations from six brain tissue samples that were assigned to us by Dr. Cohen. After finishing up our work in the lab, we then moved into working on our end of the semester presentation as well as we prepared to show the RISEbio community of faculty, students, peer mentors, parents, all other previous RISEbio students, and anyone else who wanted to join!
Research:
We were given six brain tissue samples from six different lizards by Dr. Cohen to perform the RNA isolation protocol on. This protocol is extremely time sensitive and specific in making sure we perform each step correctly. To begin we must get the brain tissue sample from the -80 degrees (Celsius) freezer, and once we have obtained the tissue, this is when it is important to work quickly. We first mass the tube and the tissue together and once we obtain that value, we put the tissue from the storage tube into a tube that contains 1 ml of QIAzol and start homogenizing the sample to break it down into the solution. It is important to work efficiently during these steps since this is when the RNA from the tissue can degrade the most and cause us to get a low concentration of RNA and no amplification when the gel is run later. Once the tissue is placed in the QIAzol and homogenized is when the degradation of the RNA becomes less of a concern if we follow the rest of the protocol correctly. We then make sure to obtain the mass of the storage tube as well so we can subtract it from the value of mass of the tube and the tissue to record the mass of the tissue alone.
Since before we got the assigned brain tissue samples, we practiced this protocol on liver tissue samples from the lizards, we had become very comfortable working through this protocol more each time we performed these techniques. The rest of the steps walk us through adding certain reagents, buffer wash solutions, and enzymes into the spin column to continue to wash out and eliminate all the other components that are within the solution, until we have the RNA isolated alone. After we have worked through the steps within the protocol, we then move forward onto checking the concentration and purity value of the isolated RNA sample to ensure we did not degrade the RNA too much which will be seen in the concentration value and that there is not any contamination of our sample which is what the purity value tells us.
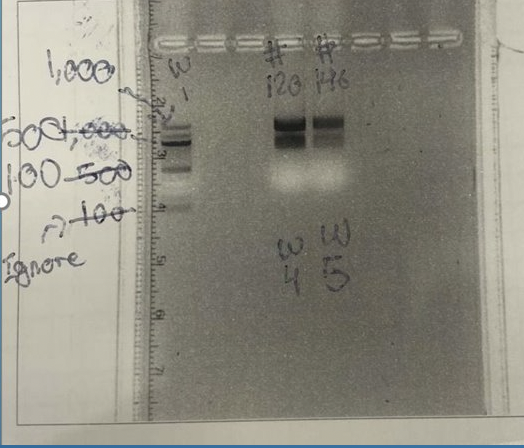
From the approximate 30 μl of isolated RNA we obtained, we take 3 μl of the sample to use to test the concentration, purity, and check for amplification by running a gel. Each time performing the protocol we were looking to obtain a concentration between 200-500 ng/μl and a purity value around 2. To check for this, we use the nanodrop machine in the lab that allows us to get these values. Once we blank the nanodrop, we use only 1 of the 3 μl to check for these values. Then after getting the values of concentration and purity recorded, we then use the rest of 2 μl we took from our isolated RNA and add gel loading dye to it and run the sample through gel electrophoresis. After the gel is finished, we take it to the gel image room so we can check for amplification. When looking for amplification we are trying to ensure that we can see the amplification of the 18S and 28S RNA bands in each sample.
We were successful in having great values of concentration, purity, and great amplification for all our six lizard brain tissue samples when isolating the RNA. This is where our research will come to a stop for now, until we come back next semester to pick back up right where we left off! To finish the semester out, we got to present our research to the RISEbio community of faculty, students, mentors, parents, and previous RISEbio students as well. We had the opportunity to hear what the other streams were up to throughout the semester and all that they had done and what they have learned so far. We got to share our specific focus on our research stream under Dr. Cohen, including getting to talk about the two genes we each selected to begin with. We described the background information we gathered on the genes cold inducible RNA binding protein (CIRBP) and hormonally upregulated neu-associated kinase (HUNK). We also went into detail about the process and experience we have gone through this past semester within the lab including all the processes, protocols, the technology we were using, and even the troubles we ran into throughout the semester. The future plans for next semester are to create cDNA from the isolated RNA we just collected and use the cDNA, along with our primers, to perform a quantitative PCR. We shared all this information with the RISEbio community and felt good about the presentation of our research afterward.
Conclusion:
Overall, this semester has been extremely rewarding with all we were able to get to do in the lab and everything we have learned from working in Dr. Cohen’s brain and behavior stream. We are appreciative of the opportunity we to get to be a part of this awesome program and the opportunities it has given to us. We are excited to continue to conduct our research and cannot wait to continue into next semester!