James Hawco and Jake Lohn
Introduction:
The majority of our time from the last post to this one has been waiting for various colonies to grow since conjugation is a process that takes place on different media for it to work. The process of conjugation is relatively simple but, as stated earlier, requires an extreme amount of time compared to any other thing we have done in our research stream.
Specific Content:
The work to perform conjugation started with our professor, Dr. Zhu, who streaked our co-transformed E. coli on LB plates and F. psychrophilum on TYES plates. The conjugation we performed was the complementation of fpsY mutant with the pKL03 plasmid. After Dr. Zhu’s preparation, we scraped the F. psychrophilum cells (fpsY mutant) off the TYES plates and suspended them in TYES media. After this, we took the F. psychrophilum cells and the E. coli cells containing pKL03 and began the next steps of conjugation. After multiple centrifugations and washes, we mixed the E. coli and F. psychrophilum together and transferred all the cells to TYES plates for the conjugation to occur. After 2 days of conjugation, we plated these mixed cells on agar media with the antibiotic erythromycin, and then incubated the plates for 7 days and checked for growth. Any of the F. psychrophilum colonies grown on the TYES erythromycin plates should have accepted the plasmid pKL03 from E. coli. We then streaked the colonies on new TYES erythromycin plates for purification. After this, we streaked a single colony on a fresh plate and incubated them until enough growth was observed.
Once enough growth had occurred, we performed a serial dilution to 10-1. We plated 100 µl of the undiluted and diluted cultures on TYES plates with erythromycin. In total, we plated 5 plates of undiluted and 5 plates of 10-1 diluted cultures. After the plating, we set the plates in the incubator for 6 days and if colonies did grow on the plates, it means the plasmid pKL03 was transferred into the mutant.
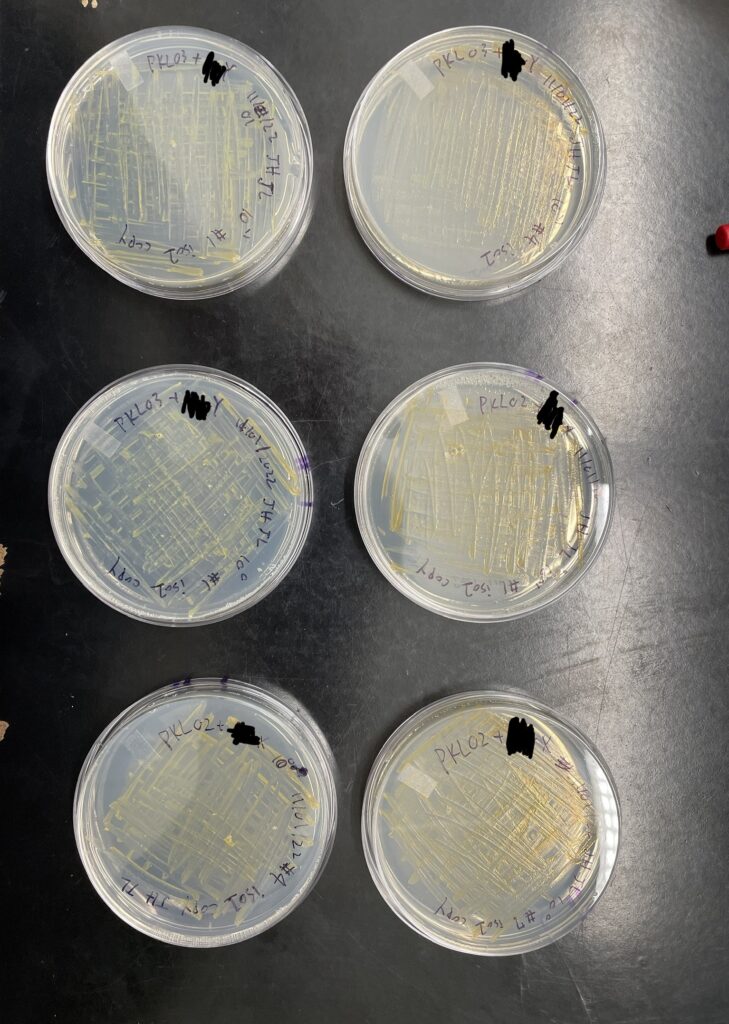
After 6 days we checked our plates and there were many colonies that had grown. We then performed streaking for the isolation of pure cultures. We picked 2 colonies from the undiluted and 2 colonies from 10-1 diluted plates. We also streaked 2 colonies on skim milk plates and casein protein plates respectively to observe the effectiveness of their protein digestion as compared to the wildtype F. psychrophilum.
Conclusions/Reflections:
Compared to before, we have only had one issue occur which made our progress not be as impacted as last time. This has allowed us to most likely be able to begin some testing with our complementation strain towards the end of the semester which would be our future plans besides finishing the end steps of conjugation.
Since this is our final blog, we would like to thank everyone who has read our blogs over the time we have been in RISEBio and we would also like to thank our research stream professor Dr. Zhu, the leading RISEBio professor Dr. Sharlin, our lab mentors Seada Sloboda, Emily Schmidtbauer, and Nick Shankey, the NSF for funding this opportunity, and MNSU for having the RISEBio program on this campus.